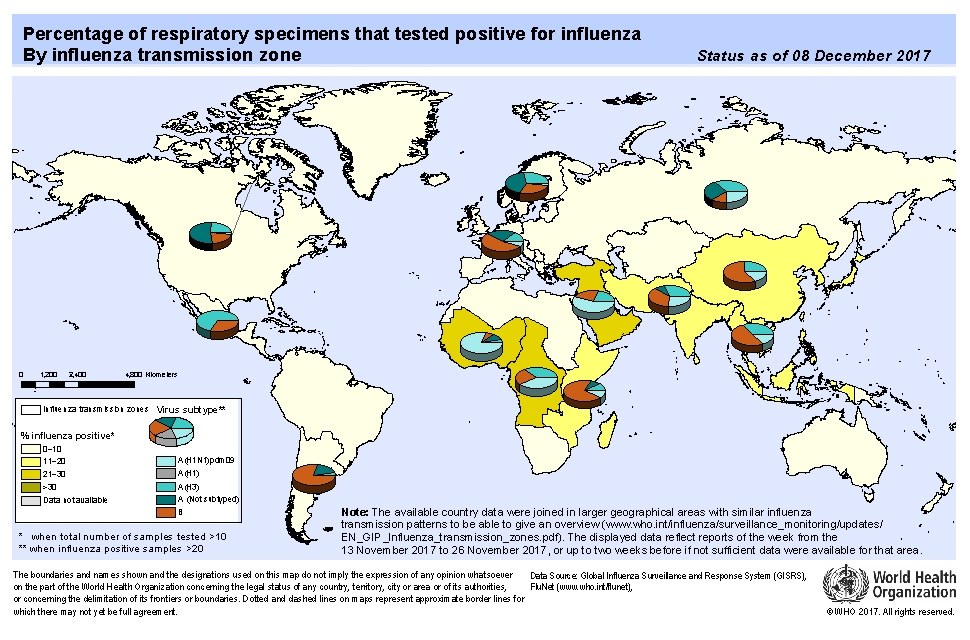
1. GLOBAL WHO SUMMARY – based on data up to Dec 08th 2017
North America: overall influenza activity continued to increase in the region, with detections of predominantly influenza A(H3N2) viruses.
Europe: influenza activity increased since the previous weeks, but remained low, with detections of predominantly influenza B viruses followed by influenza A(H3N2) viruses.
Northern Africa: sporadic influenza A virus detections were reported in Morocco and Tunisia.
Western Africa: influenza A(H1N1)pdm09 virus detections increased in Cote d’Ivoire and Ghana. In Middle Africa, influenza B detections were reported in Central African Republic.
In Eastern Africa: influenza B Yamagata-lineage virus detections were reported in Mozambique.
Western Asia: high levels of influenza activity were reported in Oman and Qatar in recent weeks, with detections of all seasonal influenza subtypes.
Central Asia: respiratory illness indicators appeared to increase in Kazakhstan and Uzbekistan in recent weeks.
Eastern Asia: influenza activity remained low in general. In Northern China, ILI and influenza percentage positive continued to increase, with influenza A(H3N2) and B Yamagata-lineage viruses predominantly detected.
2. VIROLOGICAL SURVEILLANCE AND STRAIN CHARACTERIZATION
WHO GISRS laboratories: from 13 November 2017 to 26 November 2017
NICs and other national influenza laboratories from 99 countries reported data to FluNet for the time period More than 113412 specimens were tested during that time period and 8982 were positive for influenza viruses (7,9%), of which 5617 (62.5%) were typed as influenza A and 3365 (37.5%) as influenza B. Of the sub-typed influenza A viruses, 1122 (33%) were influenza A(H1N1)pdm09 and 2273 (67%) were influenza A(H3N2). Of the characterized B viruses, 1521 (80%) belonged to the B-Yamagata lineage and 381 (20%) to the B-Victoria lineage.
NB: The recommended components for the 2017-2018 Northern hemisphere Influenza vaccine includes:
an A/Michigan/45/2015 (H1N1)pdm09-like virus; an A/Hong Kong/4801/2014 (H3N2)-like virus; a B/Brisbane/60/2008-like virus (Vic Lineage). and a B/Phuket/3073/2013-like virus (Yam Lineage) in QIV vaccine.
EUROPE weeks: Week 40 to 48/2017
While relatively few of the viruses detected in non-sentinel samples since week 40/2017 have been ascribed to a subtype or lineage, of all subtyped A viruses 82% were A(H3N2). Of influenza type B viruses ascribed to a lineage (n=76), 92% were B/Yamagata lineage and 8% were B/Victoria lineage.
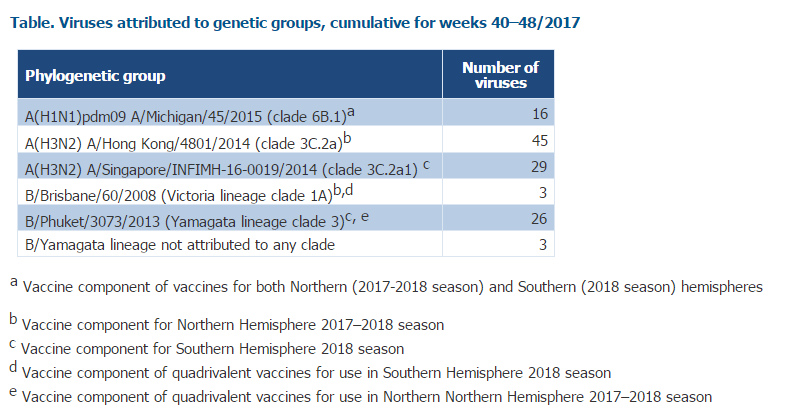
Genetic characterization of 122 viruses has been reported (see table below). Among 74 influenza A(H3N2) viruses, 45 (61%) fell in the vaccine virus component clade (3C.2a), and 29 (39%) in subclade 3C.2a1 with viruses defined by N171K, often with N121K, amino acid substitutions in the haemagglutinin. Viruses in these 2 groups are antigenically similar, but both clade and subclade are evolving rapidly with the emergence of several virus clusters defined by additional amino acid substitutions in the haemagglutinin, thereby requiring continued monitoring of antigenic characteristics. Three B/Yamagata viruses were not attributed to any clade.
See also the November report (genetic data as of week 48)
US-CDC- Until Nov 25th, 2017
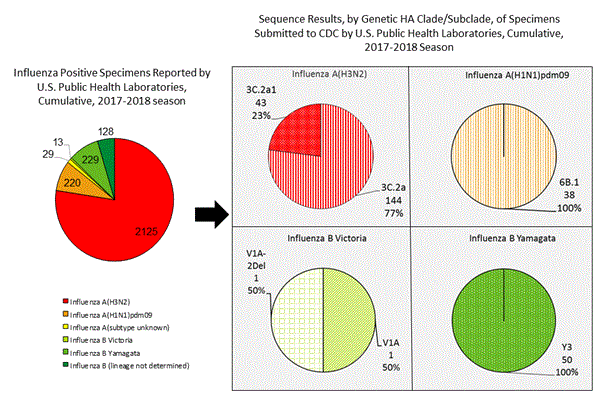
CDC has antigenically or genetically characterized 277 influenza viruses collected during October 1 – November 25, 2017 including 38 influenza A(H1N1)pdm09 viruses, 187 influenza A(H3N2) viruses, and 52 influenza B viruses.
A (H1N1)pdm09: Phylogenetic analysis of the HA genes from 38 A(H1N1)pdm09 viruses showed that all belonged to clade 6B.1. 38 A(H1N1)pdm09 viruses were antigenically characterized, and all were antigenically similar (analyzed using HI with ferret antisera) to the reference 6B.1 virus A/Michigan/45/2015, representing the recommended influenza A(H1N1)pdm09 reference virus for the 2017–18 NH influenza vaccines.
A (H3N2): Phylogenetic analysis of the HA genes from 187 A(H3N2) viruses revealed extensive genetic diversity with multiple clades/subclades co-circulating. The HA genes of circulating viruses belonged to clade 3C.2a (n=144) or subclade 3C.2a1 (n=43). 64 influenza A(H3N2) viruses were antigenically characterized, and 63 (98%) A(H3N2) viruses tested were well-inhibited (reacting at titers that were within fourfold of the homologous virus titer) by ferret antisera raised against A/Michigan/15/2014 (3C.2a), a cell propagated A/Hong Kong/4801/2014-like reference virus representing the A(H3N2) component of 2017–18 NH influenza vaccines.
B/Victoria: Phylogenetic analysis of two B/Victoria-lineage viruses indicate that all HA genes belonged to genetic clade V1A, the same genetic clade as the vaccine reference virus, B/Brisbane/60/2008. However, a small number of viruses identified in 2017 had a 6-nucleotide deletion (encoding amino acids 162 and 163) in the HA (abbreviated as V1A-2Del). One (50%) of two B/Victoria lineage viruses were well-inhibited by ferret antisera raised against cell -propagated B/Brisbane/60/2008 reference virus, representing a recommended B virus component of 2017–18 Northern Hemisphere influenza vaccines. One B/Victoria lineage virus reacted poorly (at titers that were 8-fold or greater reduced compared with the homologous virus titer) with ferret antisera raised against cell-propagated B/Brisbane/60/2008, and this virus had the two amino acid deletion in the HA of the V1A-2Del viruses.
B/Yamagata: Phylogenetic analysis of 50 influenza B/Yamagata-lineage viruses indicate that the HA genes belonged to clade Y3. A total of 14 influenza B/Yamagata-lineage viruses were antigenically characterized, and all were antigenically similar to cell propagated B/Phuket/3073/2013, the reference vaccine virus representing the influenza B/Yamagata-lineage component of the 2017–18 Northern Hemisphere QIV.
HPA-London until Week 48
The PHE Respiratory Virus Unit has characterised 88 influenza viruses detected since week 37. Of the 19 A(H1N1)pdm09 influenza viruses that have been characterised, all belong in the genetic subgroup 6B.1, which was the predominant genetic subgroup in the 2016/17 season. The four viruses antigenically analysed are similar to the A/Michigan/45/2015 Northern Hemisphere 2017/18 (H1N1)pdm09 vaccine strain.
Genetic characterisation of 50 A(H3N2) influenza viruses detected since late summer, showed that they all belong to genetic subclade 3C.2a, with 30 belonging to a cluster within this genetic subclade designated as 3C.2a1. The NH 2017/18 influenza A(H3N2) vaccine strain A/HongKong/4801/2014 belongs in genetic subclade 3C.2a.
Nineteen influenza B viruses have been analysed; 16 were characterised as belonging to the B/Yamagata/16/88-lineage and 3 belonging to the B/Victoria/2/1987-lineage. Of the influenza B viruses antigenically characterised, the B/Victoria/2/87-lineage viruses were antigenically similar to B/Brisbane/60/2008, the influenza B/Victoria-lineage component of 2017/18 NH TIV and QIV. B/Yamagata/16/88-lineage viruses were antigenically similar to B/Phuket/3073/2013, the influenza B/Yamagata-lineage component of 2016/17 NH QIV.
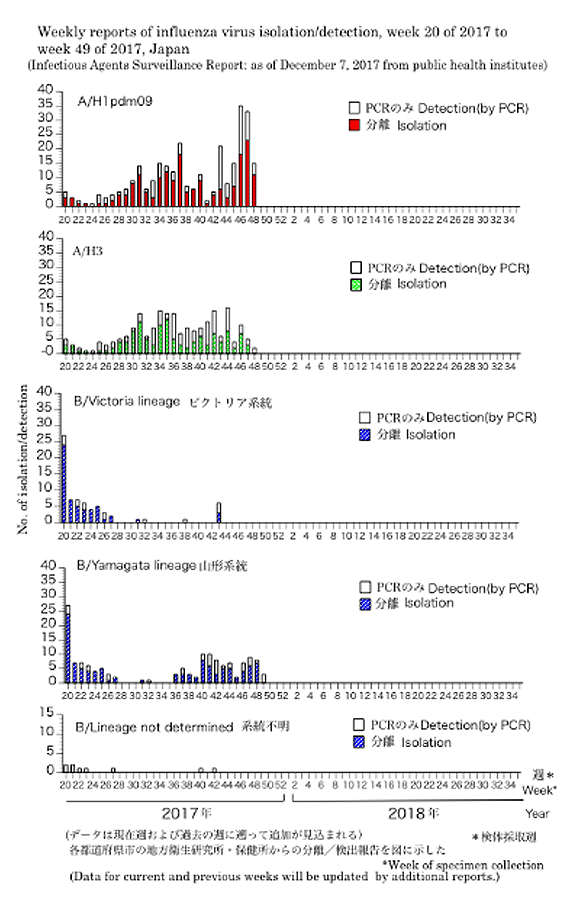
JAPAN National infectious diseases- Until Week 49
3. Real-time tracking of influenza virus evolution
Next-Flu (Sept 19th 2017 report) Neher RA, Bedford T. 2015. nextflu: real-time tracking of seasonal influenza virus evolution in humans. Bioinformatics DOI: 10.1093/bioinformatics/btv381.
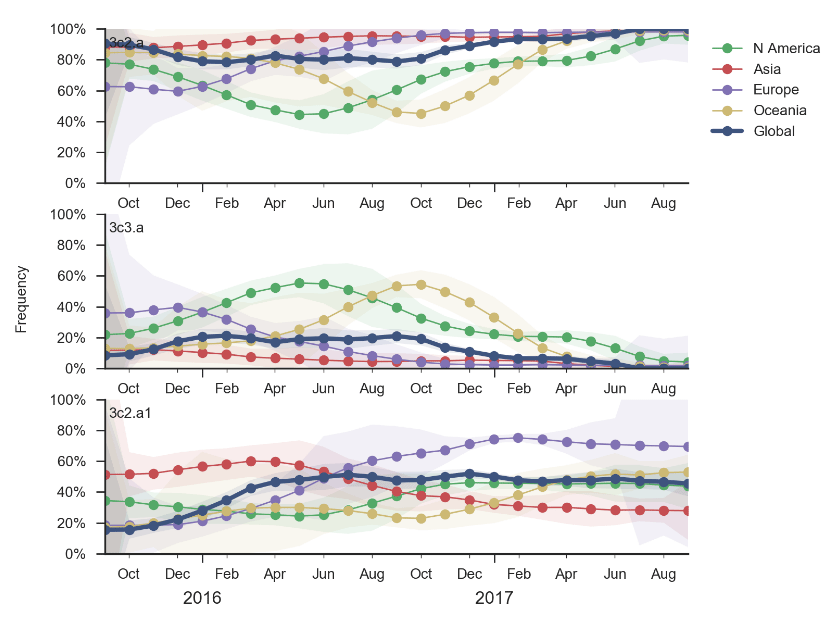
H3N2 continues to diversify with many coexisting clades, all of which carry several amino acid mutations at previously characterized epitopes sites. The majority of viruses fall into the 3c2.a clade which has been dominating globally for >3 years, but 3c3.a viruses continue to persist. The common ancestor of circulating H3N2 viruses is now more than 5 years old, which is rare for H3N2. Despite extensive genetic diversity, serological assays suggest limited, but non-zero, antigenic evolution. We expect multiple competing clades within 3c2.a to persist into the future with no clear immediate winner.
A/H1N1pdm: A clade comprising mutations S74R and I295V has risen to >60% global frequency. Although it shows no antigenic distinction by ferret HI data, the rapidity of its rise suggests a selective origin.
B/Vic: A clade with a two amino acid deletion 162-/163- has altered serological properties and is increasing in frequency, albeit slowly. Two other clades (carrying mutations K209N and V87A/I175V) have increased in frequency moderately.
B/Yam: A clade comprising M251V within clade 3 viruses continues to dominate. This is little genetic differentiation within this clade and no evidence of antigenic evolution.
4. MORE DATA FROM REGIONAL OR COUNTRIES SURVEILLANCE
- Canada
Government, FluWatch Report - China
Chinese National Influenza Center – Influenza Weekly Report-Week - Europe
ECDC/ WHO Europe Weekly Influenza update - Hong Kong
Centre for Health Protection- Flux Express Surveillance Report - PAHO
Regional Update - Russian Federation
WHO NIC at Research Institute of Influenza and D.I Ivanovsky Institute of Virology, Integrated data of influenza morbidity and diagnosis - Singapore
Ministry of Health – weekly infectious disease bulletin - Unites States Of America
Centers for Disease Control and Prevention, Weekly U.S. Influenza Surveillance Report

Этот метод сервисного обслуживания мы переняли у американских партнёров, система CORE/Обмен у них существует уже не первый десяток лет, американцы не тратят время на ремонт – они покупают готовый продукт https://akpp-moscow.ru/.
Компания Power Transmission® (Пауэр Трансмишн) – инновационный подход к ремонту автоматических коробок передач https://akpp-moscow.ru/.
Здравствуйте, сегодня рыбачили в Барабаново на спиннинг с лодки. Народу много, насчитали около 40 лодок https://fishing55.ru/.
Рассчитайте стоимость и время доставки вашего груза https://box-send.ru/.
Сэкономьте на покупке товара до 50% и получите возможность купить то, что нельзя сейчас купить в России https://box-send.ru/.
Уважаемые родители и учащиеся! По настоятельной рекомендации Роспотребнадзора классы ,в которых хотя бы один ученик имеет подтвержденный Ковид, выводятся на 7 дней, организация обучения происходит с использованием дистанционных образовательных технологий https://sch11-33.ru/.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые интегрируют непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые объединяют уникальное оформление и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые интегрируют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые интегрируют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые гармонично соединяют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые объединяют уникальное оформление и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые интегрируют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы построим сайты, которые объединяют изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые объединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые объединяют изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы создаем сайты, которые совмещают элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые гармонично соединяют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые объединяют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые сочетают в себе уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы построим сайты, которые гармонично соединяют уникальное оформление и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Отразите характер вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые гармонично соединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы создаем сайты, которые объединяют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые сочетают в себе привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Отразите характер вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые интегрируют элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют уникальное оформление и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые совмещают привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые интегрируют элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые объединяют уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые сочетают в себе привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые гармонично соединяют уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы построим сайты, которые совмещают элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые сочетают в себе уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые объединяют уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые сочетают в себе непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые совмещают изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые сочетают в себе непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые объединяют непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые интегрируют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые объединяют непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые интегрируют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют уникальное оформление и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые совмещают непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые гармонично соединяют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые сочетают в себе привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые объединяют уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые сочетают в себе непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы создаем сайты, которые совмещают изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы создаем сайты, которые совмещают изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые совмещают изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые сочетают в себе непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые гармонично соединяют непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Отразите характер вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые объединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые гармонично соединяют уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые объединяют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы построим сайты, которые объединяют привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые совмещают изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые интегрируют непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые совмещают привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые совмещают элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые сочетают в себе уникальное оформление и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые гармонично соединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые интегрируют элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы построим сайты, которые сочетают в себе привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые совмещают элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые совмещают изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые интегрируют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые гармонично соединяют элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые сочетают в себе непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые интегрируют уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые гармонично соединяют уникальное оформление и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые объединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые сочетают в себе уникальное оформление и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые объединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые объединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые сочетают в себе привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые гармонично соединяют уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы создаем сайты, которые совмещают привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые гармонично соединяют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые совмещают непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые интегрируют уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые гармонично соединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые объединяют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые совмещают элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые гармонично соединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые совмещают непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые гармонично соединяют уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые объединяют непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые интегрируют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые гармонично соединяют изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые совмещают элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые совмещают изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые совмещают уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы создаем сайты, которые совмещают уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Отразите характер вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы построим сайты, которые объединяют непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые объединяют уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые гармонично соединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Верхний город – это восемь жилых домов, квартиры в которых предлагаются с отделкой https://lucky-savills.ru/.
Boho coffee у Стокманн, ТРЦ Гринвич, 8 марта, 46, 1 этаж, рядом с бывшим Стокманн https://bohoekb.ru/.
Boho coffee у Гиперболы, ТРЦ Гринвич, 8 марта, 46, 0 этаж, напротив Гиперболы https://bohoekb.ru/.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые совмещают элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С сердечными пожеланиями, PSS.
Электромеханические защелки для дверей незаменимы там, где наблюдается очень интенсивный поток входящих и выходящих людей. Для аварийных выходов такие устройства также идеально подходят https://adamsrite.ru/.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые сочетают в себе уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Наши резиденты являются участниками популярных телепроектов: Comedy Club, Comedy Баттл, Не Спать, Убойная Лига, Смех Без Правил и многие другие https://eventcomedy.ru/!
При подаче напряжения фиксатор электрозащелки разблокируется и дверь может быть открыта. Когда же дверь захлопывается, ригель защелкивается, и дверь оказывается надежно заперта https://adamsrite.ru/.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые совмещают непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Наши артисты имеют опыт ведения мероприятий любого уровня и масштаба https://eventcomedy.ru/.
Если есть необходимость в том, чтобы дверь в течение долгого времени была открыта, то можно выбрать длительную разблокировку https://adamsrite.ru/.
Но гораздо удобней использовать функцию электрического открывания электромеханической защелки https://adamsrite.ru/.
Пригласите резидентов Comedy Club Saint-Petersburg к себе на праздник! Наши артисты с удовольствием выступят на Вашем мероприятии с юмористическими номерами, сделав это событие запоминающимся https://eventcomedy.ru/!
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые гармонично соединяют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Наши артисты проведут мероприятие любого уровня: свадьба, корпоративный вечер, презентация, день рождения или празднование Нового года https://eventcomedy.ru/!
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые интегрируют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С лучшими пожеланиями, PSS.
Мы провели более 3000 мероприятий, обретя бесценный опыт создания и реализации уникальных концепций https://eventcomedy.ru/.
Дверь, оборудованная защелкой такого типа, оснащена снаружи и изнутри неподвижной ручкой, никак не связанной с замком и всегда открывается ключом https://adamsrite.ru/.
Представьте себе сайт, который сразу же запоминается, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые совмещают элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые сочетают в себе изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые интегрируют привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые гармонично соединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые совмещают непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые сочетают в себе непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые интегрируют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые объединяют элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые объединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые объединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые объединяют уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С сердечными пожеланиями, PSS.
Мы собрали самые достойные решения и с радостью поделимся с Вами https://skbbank-kredit.ru/.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые объединяют уникальное оформление и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Обсуждайте и задавайте ваши вопросы, мы всегда поможем вам https://skbbank-kredit.ru/!
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
На сайте есть много необычных решений и хитростей от наших профи https://skbbank-kredit.ru/.
Мы убеждены, что кухня — это сердце дома. Каждая наша кухня спроектирована с вниманием к мелочам и заботой о вас https://kuppersberg-deluxe.ru/.
Купить кухню от производителя по выгодной цене https://kuppersberg-deluxe.ru/.
Кухни на заказ в Москве https://kuppersberg-deluxe.ru/.
Дизайнер приедет к вам, выполнит замеры и предложит предварительные варианты проекта https://kuppersberg-deluxe.ru/.
Наш опыт позволяет учесть все нюансы планировки и размеров вашего дома и изготовить кухню, которая будет радовать вас каждый день красотой и функциональностью https://kuppersberg-deluxe.ru/.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые интегрируют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые интегрируют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Отразите характер вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые интегрируют элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые совмещают элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые объединяют уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые совмещают уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые совмещают привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые объединяют уникальное оформление и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые объединяют уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы построим сайты, которые интегрируют уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы построим сайты, которые сочетают в себе привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы построим сайты, которые сочетают в себе привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые объединяют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые совмещают уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы построим сайты, которые интегрируют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые сочетают в себе изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые интегрируют элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые объединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые совмещают уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые сочетают в себе уникальное оформление и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые интегрируют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые объединяют элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые интегрируют уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда рады помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые интегрируют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы создаем сайты, которые гармонично соединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые сочетают в себе уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые интегрируют элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые совмещают изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые совмещают непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы построим сайты, которые объединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые интегрируют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые гармонично соединяют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые объединяют элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые гармонично соединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые сочетают в себе изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые интегрируют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые совмещают непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы создаем сайты, которые гармонично соединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы построим сайты, которые совмещают элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые сочетают в себе элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы создаем сайты, которые совмещают непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые совмещают привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые совмещают уникальное оформление и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые гармонично соединяют уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые совмещают непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые гармонично соединяют уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые гармонично соединяют непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые объединяют элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые объединяют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые совмещают привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые гармонично соединяют уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые объединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые интегрируют уникальное оформление и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые объединяют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые сочетают в себе непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые совмещают непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые сочетают в себе уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые интегрируют уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые объединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые интегрируют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы построим сайты, которые интегрируют непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы построим сайты, которые интегрируют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые интегрируют изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые сочетают в себе элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые интегрируют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые объединяют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые объединяют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые сочетают в себе элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы построим сайты, которые объединяют уникальное оформление и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Посетите нас на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые совмещают элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые совмещают изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые сочетают в себе уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые совмещают непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые гармонично соединяют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые интегрируют непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые совмещают уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые совмещают непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые интегрируют непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Отразите характер вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы создаем сайты, которые совмещают изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые совмещают непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые сочетают в себе привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые сочетают в себе привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые объединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые интегрируют уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые объединяют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые совмещают элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые объединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые гармонично соединяют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые совмещают изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые интегрируют изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы построим сайты, которые сочетают в себе уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые интегрируют непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые объединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы построим сайты, которые сочетают в себе элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые объединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые совмещают элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые совмещают привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Отразите характер вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы построим сайты, которые интегрируют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы проектируем сайты, которые интегрируют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы построим сайты, которые интегрируют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы создаем сайты, которые интегрируют уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые совмещают уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые совмещают элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые сочетают в себе изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые гармонично соединяют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые совмещают элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые интегрируют уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые сочетают в себе непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые интегрируют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы построим сайты, которые интегрируют уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы построим сайты, которые объединяют элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые интегрируют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые сочетают в себе привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы создаем сайты, которые интегрируют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые совмещают изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Посетите нас на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые объединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые совмещают уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые совмещают изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые объединяют уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые объединяют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые совмещают уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые совмещают элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда готовы помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые сочетают в себе изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые объединяют изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые сочетают в себе непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые гармонично соединяют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые сочетают в себе элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые совмещают изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые сочетают в себе элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые интегрируют изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы всегда готовы помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые сочетают в себе элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые объединяют непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы построим сайты, которые совмещают уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые сочетают в себе изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые интегрируют элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые объединяют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые гармонично соединяют элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые совмещают непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые объединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы создаем сайты, которые совмещают непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые совмещают непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые интегрируют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые объединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые совмещают привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые объединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые объединяют привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы построим сайты, которые интегрируют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и обеспечивает эффективность. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые гармонично соединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые интегрируют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые объединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы создаем сайты, которые объединяют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые гармонично соединяют непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Идеальное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы построим сайты, которые совмещают уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы создаем сайты, которые сочетают в себе непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые объединяют непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые объединяют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые гармонично соединяют уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые объединяют непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые объединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы построим сайты, которые сочетают в себе непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые совмещают уникальное оформление и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые гармонично соединяют непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые сочетают в себе уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые интегрируют изысканный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Отзывчивый дизайн: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы создаем сайты, которые сочетают в себе уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе уникальное оформление и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые объединяют привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые совмещают привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые гармонично соединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые объединяют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые сочетают в себе уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые объединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые объединяют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые интегрируют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Отразите характер вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– Оптимизация под поисковики: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые сочетают в себе непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые совмещают изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые сочетают в себе элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда готовы помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые гармонично соединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые совмещают непревзойденный стиль и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые совмещают изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы разрабатываем сайты, которые объединяют изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые объединяют элегантный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые объединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые совмещают уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы построим сайты, которые гармонично соединяют непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и обеспечивает эффективность. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация под поисковики: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на всех этапах.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые сочетают в себе уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и обеспечивает эффективность. В PSS-Studio мы создаем сайты, которые объединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы построим сайты, которые сочетают в себе уникальное оформление и надежную работу, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые объединяют привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые совмещают элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Эксклюзивный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы всегда на связи помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые совмещают привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Повышение видимости в поиске: Повысьте ваш рейтинг в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и гарантирует успех. В PSS-Studio мы проектируем сайты, которые сочетают в себе непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит выбрать нас?
– Особенный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы всегда готовы помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и гарантирует успех. В PSS-Studio мы построим сайты, которые сочетают в себе уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Универсальное представление: Оптимальный просмотр на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые совмещают привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и приносит результаты. В PSS-Studio мы построим сайты, которые совмещают непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Мобильная адаптация: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые сочетают в себе изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые сочетают в себе элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к непревзойденному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые гармонично соединяют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые сочетают в себе привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на всех этапах.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы отправиться в путешествие к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые совмещают непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Выделите особенности вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые совмещают элегантный дизайн и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– SEO-оптимизация: Повысьте ваш рейтинг в результатах поиска.
– Всегда на связи: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Приходите к нам на https://pss-studio.ru/, чтобы пройти путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые интегрируют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Мобильная адаптация: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Готовность помочь в любой момент: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые совмещают непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые сочетают в себе привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Безупречный вид на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Приходите к нам на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые совмещают уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Приглашаем на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие прямо сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы создаем сайты, которые интегрируют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют привлекательный внешний вид и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к выдающемуся сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые гармонично соединяют уникальное оформление и отличную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на каждом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие прямо сейчас. Посетите нас на https://pss-studio.ru/, чтобы начать путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы разрабатываем сайты, которые интегрируют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Мобильная адаптация: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда готовы помочь вам на каждой ступени.
Специальное предложение: Воспользуйтесь бесплатной консультацией и уникальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые объединяют изысканный дизайн и высокую производительность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Превосходное отображение на любом устройстве.
– Оптимизация для поисковых систем: Увеличьте вашу видимость в результатах поиска.
– Постоянная поддержка: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые объединяют непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Уникальный дизайн: Подкрепите идентичность вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют привлекательный внешний вид и надежную работу, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит выбрать нас?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Измените ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые сочетают в себе уникальное оформление и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Запишитесь на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы проектируем сайты, которые объединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Отразите характер вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– Оптимизация для поисковых систем: Повысьте ваш рейтинг в результатах поиска.
– Готовность помочь в любой момент: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Обновите ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы выступить на путь к замечательному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые объединяют непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит довериться нам?
– Эксклюзивный дизайн: Отразите характер вашего бренда.
– Гибкость отображения: Безупречный вид на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые совмещают уникальное оформление и безупречную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– SEO-оптимизация: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к непревзойденному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы построим сайты, которые гармонично соединяют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация под поисковики: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Запишитесь на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы начать ваше путешествие к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые гармонично соединяют непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Персонализированный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Постоянная поддержка: Мы всегда к вашим услугам помочь вам на любой стадии.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать путь к исключительному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые объединяют привлекательный внешний вид и отличную функциональность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Запишитесь на бесплатную консультацию и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и помогает достигнуть целей. В PSS-Studio мы создаем сайты, которые объединяют элегантный дизайн и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые сочетают в себе непревзойденный стиль и высокую производительность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Отразите характер вашего бренда.
– Мобильная адаптация: Превосходное отображение на любом устройстве.
– Улучшение позиций в поисковых системах: Расширьте ваше присутствие в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Обновите ваше цифровое присутствие сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать путь к непревзойденному сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы создаем сайты, которые интегрируют элегантный дизайн и безупречную функциональность, гарантируя, что ваш бренд будет заметен в Интернете.
Почему стоит довериться нам?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда готовы помочь вам на любом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к впечатляющему сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и помогает достигнуть целей. В PSS-Studio мы построим сайты, которые сочетают в себе элегантный дизайн и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит предпочесть нас?
– Индивидуальный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– SEO-оптимизация: Укрепите вашу позицию в результатах поиска.
– Всегда на связи: Мы всегда рады помочь вам на всех этапах.
Специальное предложение: Воспользуйтесь бесплатной консультацией и специальную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы выступить на путь к исключительному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы проектируем сайты, которые совмещают непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Персонализированный дизайн: Подчеркните уникальность вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– SEO-оптимизация: Увеличьте вашу видимость в результатах поиска.
– Готовность помочь в любой момент: Мы постоянно доступны помочь вам на всех этапах.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и способствует вашему росту. В PSS-Studio мы построим сайты, которые сочетают в себе изысканный дизайн и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит предпочесть нас?
– Эксклюзивный дизайн: Усилите индивидуальность вашего бренда.
– Универсальное представление: Безупречный вид на любом устройстве.
– SEO-оптимизация: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Получите бесплатную консультацию и эксклюзивную скидку на наши услуги по разработке сайтов, если свяжетесь с нами в течение следующих 48 часов.
Преобразите ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы начать путь к выдающемуся сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и приносит результаты. В PSS-Studio мы построим сайты, которые гармонично соединяют изысканный дизайн и отличную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Безупречный вид на любом устройстве.
– Улучшение позиций в поисковых системах: Повысьте ваш рейтинг в результатах поиска.
– Непрерывная помощь: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые интегрируют изысканный дизайн и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Превосходное отображение на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Всегда на связи: Мы всегда к вашим услугам помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы выступить на путь к впечатляющему сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который сразу же запоминается, но и помогает достигнуть целей. В PSS-Studio мы проектируем сайты, которые объединяют привлекательный внешний вид и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит рассмотреть наше предложение?
– Особенный дизайн: Усилите индивидуальность вашего бренда.
– Адаптивность: Идеальное отображение на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Круглосуточная поддержка: Мы всегда рады помочь вам на каждом этапе.
Специальное предложение: Получите бесплатную консультацию и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Обновите ваше цифровое присутствие сейчас. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы начать ваше путешествие к исключительному сайту.
С сердечными пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые интегрируют уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет бросаться в глаза в Интернете.
Почему стоит рассмотреть наше предложение?
– Индивидуальный дизайн: Подчеркните уникальность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Улучшение позиций в поисковых системах: Укрепите вашу позицию в результатах поиска.
– Готовность помочь в любой момент: Мы всегда готовы помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и эксклюзивную скидку на наши услуги по разработке сайтов, если дадите обратную связь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к впечатляющему сайту.
С наилучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и приносит результаты. В PSS-Studio мы разрабатываем сайты, которые интегрируют непревзойденный стиль и безупречную функциональность, гарантируя, что ваш бренд будет оставаться незабываемым в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Отразите характер вашего бренда.
– Отзывчивый дизайн: Великолепное представление на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Круглосуточная поддержка: Мы постоянно доступны помочь вам на каждом этапе.
Специальное предложение: Используйте возможность бесплатной консультации и особую скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие сегодня. Приходите к нам на https://pss-studio.ru/, чтобы начать ваше путешествие к непревзойденному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и помогает достигнуть целей. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют уникальное оформление и превосходную эффективность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит обратиться к нам?
– Особенный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Заставьте заметить ваш бренд в результатах поиска.
– Круглосуточная поддержка: Мы всегда к вашим услугам помочь вам на любом этапе.
Специальное предложение: Воспользуйтесь бесплатной консультацией и особую скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Преобразите ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы пройти путь к замечательному сайту.
С искренними пожеланиями, PSS.
Представьте себе сайт, который не только привлекает внимание, но и обеспечивает эффективность. В PSS-Studio мы построим сайты, которые сочетают в себе непревзойденный стиль и надежную работу, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Гибкость отображения: Оптимальный просмотр на любом устройстве.
– Улучшение позиций в поисковых системах: Увеличьте вашу видимость в результатах поиска.
– Всегда на связи: Мы постоянно доступны помочь вам на любом этапе.
Специальное предложение: Получите бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Трансформируйте ваше цифровое присутствие немедленно. Посетите нас на https://pss-studio.ru/, чтобы отправиться в путешествие к исключительному сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который захватывает дух, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые интегрируют уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит выбрать нас?
– Персонализированный дизайн: Подкрепите идентичность вашего бренда.
– Адаптивность: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Постоянная поддержка: Мы всегда на связи помочь вам на каждой ступени.
Специальное предложение: Используйте возможность бесплатной консультации и дополнительную скидку на наши услуги по разработке сайтов, если напишете нам в течение следующих 48 часов.
Измените ваше цифровое присутствие немедленно. Зайдите на наш сайт на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и приносит результаты. В PSS-Studio мы воплощаем в жизнь сайты, которые гармонично соединяют непревзойденный стиль и превосходную эффективность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит обратиться к нам?
– Индивидуальный дизайн: Выделите особенности вашего бренда.
– Адаптивность: Оптимальный просмотр на любом устройстве.
– Оптимизация под поисковики: Заставьте заметить ваш бренд в результатах поиска.
– Непрерывная помощь: Мы постоянно доступны помочь вам на любой стадии.
Специальное предложение: Используйте возможность бесплатной консультации и специальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Измените ваше цифровое присутствие уже сегодня. Посетите нас на https://pss-studio.ru/, чтобы начать ваше путешествие к замечательному сайту.
С лучшими пожеланиями, PSS.
Представьте себе сайт, который притягивает взгляды, но и способствует вашему росту. В PSS-Studio мы разрабатываем сайты, которые объединяют привлекательный внешний вид и высокую производительность, гарантируя, что ваш бренд будет выделяться в Интернете.
Почему стоит довериться нам?
– Уникальный дизайн: Усилите индивидуальность вашего бренда.
– Отзывчивый дизайн: Оптимальный просмотр на любом устройстве.
– Повышение видимости в поиске: Увеличьте вашу видимость в результатах поиска.
– Непрерывная помощь: Мы всегда на связи помочь вам на любой стадии.
Специальное предложение: Приглашаем на бесплатную консультацию и дополнительную скидку на наши услуги по разработке сайтов, если откликнитесь в течение следующих 48 часов.
Преобразите ваше цифровое присутствие сейчас. Приглашаем вас на наш сайт на https://pss-studio.ru/, чтобы пройти путь к впечатляющему сайту.
С теплыми пожеланиями, PSS.
Представьте себе сайт, который вызывает интерес, но и гарантирует успех. В PSS-Studio мы создаем сайты, которые сочетают в себе уникальное оформление и высокую производительность, гарантируя, что ваш бренд будет привлекать внимание в Интернете.
Почему стоит предпочесть нас?
– Уникальный дизайн: Подчеркните уникальность вашего бренда.
– Универсальное представление: Великолепное представление на любом устройстве.
– Оптимизация для поисковых систем: Расширьте ваше присутствие в результатах поиска.
– Готовность помочь в любой момент: Мы всегда рады помочь вам на каждой ступени.
Специальное предложение: Приглашаем на бесплатную консультацию и уникальную скидку на наши услуги по разработке сайтов, если ответите в течение следующих 48 часов.
Перевоплотите ваше цифровое присутствие уже сегодня. Ознакомьтесь с нашим предложением на https://pss-studio.ru/, чтобы отправиться в путешествие к выдающемуся сайту.
С наилучшими пожеланиями, PSS.
Невероятное качество баз! Рекомендую всем, кто хочет быть на пике https://bazydlyaxrumerkupitt.ru/!
Мы работаем над увеличением скорости и совместимости с новыми платформами для вашего удобства https://bazydlyaxrumerkupitt.ru/.
Наши базы – это не просто данные, это ключ к доминированию в поисковых системах. Доверьте свой рост нам https://bazydlyaxrumerkupitt.ru/!
Покорите вершины рейтингов вместе с нами https://bazydlyaxrumerkupitt.ru/!
Присоединяйтесь к нам и узнайте секреты итальянской кухни от наших шеф-поваров https://kupitkuhnyucena6.ru/!
Погрузитесь в мир волшебства и вкуса вместе с нашими талантливыми поварами и артистами https://kupitkuhnyucena6.ru/!
Доверьте нам создание кухни вашей мечты и погрузитесь в мир вкуса и уюта https://kupitkuhnyucena6.ru/!
Участвуйте в увлекательных мастер-классах и соревнованиях, чтобы стать настоящим шеф-поваром https://kupitkuhnyucena6.ru/!
Кухни от КухниМаркет – это просто сказка! Каждый день – праздник https://kupitkuhnyumagazin.ru/!
Мы – команда мастеров, создающих кухни мечты для каждого клиента. Вдохновляйтесь идеями и воплощайте их в жизнь с нами https://kupitkuhnyumagazin.ru/!
Забудьте о скучных кухнях! Наши дизайнеры помогут воплотить ваши самые дерзкие фантазии в реальность. Выберите стиль, цвет и форму – и ваша кухня станет настоящим произведением искусства https://kupitkuhnyumagazin.ru/!
Наша миссия – сделать кухню не просто местом приготовления пищи, а настоящим источником радости и вдохновения https://kupitkuhnyumagazin.ru/.
Лучший выбор для тех, кто ценит качество и стиль. Рекомендую всем https://kupitkuhnyumagazin.ru/!
Кухни просто супер! Не могу нарадоваться своей новой кухне, спасибо фабрике https://kupitkuhnyunedorogo.ru/!
Лучшее решение для кухни! Качество на высоте, заказал здесь и не пожалел https://kupitkuhnyunedorogo.ru/.
Откройте для себя новые вкусы и ароматы нашего мирового меню в уютной атмосфере https://kupitkuhnyucena6.ru/!
Мечтаете о кухне вашей мечты? Мы воплотим ваши желания https://kupitkuhnyumagazin.ru/!
Мы – команда профессионалов, которая превращает ваши мечты в реальность. Каждая кухня у нас – это произведение искусства и функциональности https://kupitkuhnyunedorogo.ru/.
Мы не просто продаём кухни, мы создаём шедевры, которые будут восхищать вас каждый день https://kupitkuhnyunedorogo.ru/.
Мы используем только лучшие материалы и технологии для создания кухонь, которые прослужат вам долгие годы https://kupitkuhnyu-ot-proizvoditelya.ru/.
Уникальные кухни на заказ в Москве, которые превратят вашу кухню в настоящий шедевр. Доверьте свои кулинарные фантазии нам https://kupitkuhnyu-ot-proizvoditelya.ru/!
Индивидуальные дизайны кухонь, которые вдохнут жизнь в ваш дом https://kupitkuhnyu-ot-proizvoditelya.ru/.
Наша страсть к дизайну и индивидуальному подходу позволяет нам создавать кухни, которые отражают ваш стиль и вкус https://kupitkuhnyu-ot-proizvoditelya.ru/.
Мы – команда профессионалов, готовых ускорить ваш путь к онлайн-успеху с помощью уникальных баз для XRumer и GSA Search Engine Ranker https://bazydlyaxrumerkupitt.ru/.
С нами ваш сайт станет непобедимым в борьбе за первые места в поисковой выдаче https://bazydlyaxrumerkupitt.ru/.
Каждая кухня уникальна и создана специально для вас, отражая вашу индивидуальность и стиль https://kupitkuhnyu-ot-proizvoditelya.ru/.
Наша команда опытных специалистов обеспечит качественный и профессиональный монтаж вашей кухни https://kupitkuhnyu-ot-proizvoditelya.ru/.
mexican online pharmacies prescription drugs: mexican pharmacy – mexican mail order pharmacies
online pharmacy india http://indiaph24.store/# top 10 pharmacies in india
top 10 pharmacies in india
Мы – команда страстных кулинаров, которые превращают ваши мечты о кухне в реальность https://kupitkuhnyucena6.ru/.
great article
That is really interesting, You’re a very professional blogger. I have joined your rss feed and sit up for in the hunt for extra of your magnificent post. Additionally, I’ve shared your website in my social networks!
Циклёвка паркета: особенности и этапы услуги
Циклёвка паркета — это процесс восстановления внешнего вида паркетного пола путём удаления верхнего повреждённого слоя и возвращения ему первоначального вида. Услуга включает в себя несколько этапов:
Подготовка: перед началом работы необходимо защитить мебель и другие предметы от пыли и грязи, а также удалить плинтусы.
Шлифовка: с помощью шлифовальной машины удаляется старый лак и верхний повреждённый слой древесины.
Шпатлёвка: после шлифовки поверхность паркета шпатлюется для заполнения трещин и выравнивания поверхности.
Грунтовка: перед нанесением лака паркет грунтуется для улучшения адгезии и защиты от плесени и грибка.
Нанесение лака: лак наносится в несколько слоёв с промежуточной шлифовкой между ними.
Полировка: после нанесения последнего слоя лака паркет полируется для придания поверхности блеска и гладкости.
Циклёвка паркета позволяет обновить внешний вид пола, восстановить его структуру и продлить срок службы.
Сайт: ykladka-parketa.ru Циклёвка паркета
Лендинг-пейдж — это одностраничный сайт, предназначенный для рекламы и продажи товаров или услуг, а также для сбора контактных данных потенциальных клиентов. Вот несколько причин, почему лендинг-пейдж важен для бизнеса:
Увеличение узнаваемости компании. Лендинг-пейдж позволяет представить компанию и её продукты или услуги в выгодном свете, что способствует росту узнаваемости бренда.
Повышение продаж. Заказать лендинг можно здесь – 1landingpage.ru Одностраничные сайты позволяют сосредоточиться на конкретных предложениях и акциях, что повышает вероятность совершения покупки.
Оптимизация SEO-показателей. Лендинг-пейдж создаются с учётом ключевых слов и фраз, что улучшает позиции сайта в результатах поиска и привлекает больше целевых посетителей.
Привлечение новой аудитории. Одностраничные сайты могут использоваться для продвижения новых продуктов или услуг, а также для привлечения внимания к определённым кампаниям или акциям.
Расширение клиентской базы. Лендинг-пейдж собирают контактные данные потенциальных клиентов, что позволяет компании поддерживать связь с ними и предлагать дополнительные услуги или товары.
Простота генерации лидов. Лендинг-пейдж предоставляют краткую и понятную информацию о продуктах или услугах, что облегчает процесс принятия решения для потенциальных клиентов.
Сбор персональных данных. Лендинг-пейдж позволяют собирать информацию о потенциальных клиентах, такую как email-адрес, имя и контактные данные, что помогает компании лучше понимать свою аудиторию и предоставлять более персонализированные услуги.
Улучшение поискового трафика. Лендинг-пейдж создаются с учётом определённых поисковых запросов, что позволяет привлекать больше целевых посетителей на сайт.
Эффективное продвижение новой продукции. Лендинг-пейдж можно использовать для продвижения новых товаров или услуг, что позволяет привлечь внимание потенциальных клиентов и стимулировать их к покупке.
Лёгкий процесс принятия решений. Лендинг-пейдж содержат только самую необходимую информацию, что упрощает процесс принятия решения для потенциальных клиентов.
В целом, лендинг-пейдж являются мощным инструментом для продвижения бизнеса, увеличения продаж и привлечения новых клиентов.
заказать лендинг пейдж
Thanks for your posting. Another point is that being photographer involves not only problems in recording award-winning photographs but also hardships in acquiring the best photographic camera suited to your requirements and most especially struggles in maintaining the grade of your camera. This can be very accurate and visible for those photography enthusiasts that are in to capturing the particular nature’s captivating scenes : the mountains, the particular forests, the actual wild or even the seas. Going to these amazing places unquestionably requires a digicam that can surpass the wild’s hard settings.
Hey! Someone in my Facebook group shared this site with us so I came to give it a look. I’m definitely loving the information. I’m book-marking and will be tweeting this to my followers! Outstanding blog and terrific style and design.
Excellent site. Lots of useful info here. I?m sending it to several friends ans also sharing in delicious. And naturally, thanks to your sweat!
Does your website have a contact page? I’m having trouble locating it but, I’d like to shoot you an email. I’ve got some creative ideas for your blog you might be interested in hearing. Either way, great website and I look forward to seeing it expand over time.
สล็อต PG เว็บตรง
Terrific work! That is the type of information that are meant to be shared across the net. Disgrace on the seek engines for now not positioning this put up upper! Come on over and visit my website . Thank you =)
I?m not sure where you are getting your info, but great topic. I needs to spend some time learning much more or understanding more. Thanks for great information I was looking for this info for my mission.
pin-up 141: pin up 360 – pin up kazino
I have noticed that costs for online degree pros tend to be a terrific value. Like a full 4-year college Degree in Communication from The University of Phoenix Online consists of Sixty credits with $515/credit or $30,900. Also American Intercontinental University Online offers a Bachelors of Business Administration with a entire course element of 180 units and a tariff of $30,560. Online degree learning has made having your college diploma been so cool because you may earn your own degree through the comfort of your house and when you finish from office. Thanks for all the tips I’ve learned from your web site.
pin up giris https://azerbaijancuisine.com/# pin up 306
pin-up online casino
Fantastic site. Lots of useful info here. I?¦m sending it to some buddies ans additionally sharing in delicious. And certainly, thanks on your effort!
Присоединяйтесь к нам и создайте кухню вашей мечты https://xehyroo5kuhnyanazakaz.ru/!
Превратите свою кухню в шедевр https://xehyroo5kuhnyanazakaz.ru/!
Мы – команда страстных кулинаров и дизайнеров, создающая кухни мечты для вас https://tovudyi4kuhnyanazakaz.ru/!
Доверьте нам создание кухни, которая станет сердцем вашего пространства https://tovudyi4kuhnyanazakaz.ru/.
Level up your game with insider tips and tricks for serious punters https://cricetc1xbetr1xbetcc2.ru/.
Присоединяйтесь к нам и откройте новый уровень кулинарного опыта https://guzywia4kuhnyanazakaz.ru/!
Наши кухни не просто мебель, это произведения искусства, наполненные вдохновением и инновациями https://guzywia4kuhnyanazakaz.ru/.
У нас нет предела совершенству! Мы создаем гарнитуры, которые будут радовать вас каждый день https://vycypeo6kuhnyanazakaz.ru/.
Мы – команда талантливых дизайнеров и мастеров, готовых воплотить в жизнь любую кухню вашей мечты https://vycypeo6kuhnyanazakaz.ru/.
Welcome to the ultimate destination for cricket enthusiasts who want to spice up the game with some adrenaline-pumping bets https://1xbetecricete1xbetei4.ru/!
At CricketBet, we bring the excitement of cricket betting to the vibrant landscape of the United Arab Emirates. Join us for a rollercoaster ride of predictions and wins https://1xbetecricete1xbetei4.ru/!
Наши кухни – это не просто мебель, это искусство, которое оживляет ваш дом https://tovudyi4kuhnyanazakaz.ru/.
Мы – команда талантливых дизайнеров, воплощаем в жизнь ваши самые дерзкие кулинарные фантазии https://xehyroo5kuhnyanazakaz.ru/.
Welcome to the ultimate destination for cricket enthusiasts who want to spice up the game with some adrenaline-pumping bets https://1xbeticricetc1xbetti5.ru/!
At CricketBet, we bring the excitement of cricket betting to the vibrant landscape of the United Arab Emirates. Join us for a rollercoaster ride of predictions and wins https://1xbeticricetc1xbetti5.ru/!
Уникальные кухонные шедевры ждут вас https://vycypeo6kuhnyanazakaz.ru/!
Мы – команда страстных энтузиастов, создающих кухни, которые превращают готовку в волшебство https://guzywia4kuhnyanazakaz.ru/.
Level up your game with insider tips and tricks for serious punters https://1xbett1xbetc1xbetir4.ru/.
У нас вы найдете самые необычные идеи для вашей кухни, которые заставят вас влюбиться в готовку снова https://kitubeu2kuhnyanazakaz.ru/!
Experience the thrill of betting on cricket matches in the heart of the United Arab Emirates. Get ready to win big and cheer for your favorite teams https://1xbett1xbetc1xbetir4.ru/!
Погрузитесь в мир кулинарных искусств, где каждая кухня – это отражение стиля и вкуса https://kitubeu2kuhnyanazakaz.ru/.
https://northern-doctors.org/# mexican pharmacy
buying from online mexican pharmacy: mexican northern doctors – п»їbest mexican online pharmacies
Откройте для себя новые гастрономические горизонты и дизайнерские возможности https://kitubeu2kuhnyanazakaz.ru/.
Наши кухни – это не просто мебель, это произведения искусства, которые вдохновляют на кулинарные подвиги https://xehyroo5kuhnyanazakaz.ru/!
Our team is as passionate about cricket as you are, and we’re here to make your betting experience unforgettable https://cricetc1xbeticricetec2.ru/.
https://northern-doctors.org/# mexico pharmacy
mexican online pharmacies prescription drugs: mexican pharmacy online – mexico pharmacy
Welcome to the ultimate destination for cricket enthusiasts who want to spice up the game with some adrenaline-pumping bets https://cricetc1xbeticricetec2.ru/!
mexican border pharmacies shipping to usa northern doctors pharmacy mexican drugstore online
Откройте мир волшебства в каждой кухне от фабрики КухниФаб. Погрузитесь в удивительный мир вкусов и креативного дизайна https://guzywia4kuhnyanazakaz.ru/!
Thanks , I have recently been searching for information about this subject for ages and yours is the best I have discovered so far. But, what about the conclusion? Are you sure about the source?
mexico pharmacy: northern doctors – medication from mexico pharmacy
п»їbest mexican online pharmacies: Mexico pharmacy that ship to usa – mexico drug stores pharmacies
Our team is as passionate about cricket as you are, and we’re here to make your betting experience unforgettable https://cricetc1xbetr1xbetcc2.ru/.
https://northern-doctors.org/# buying prescription drugs in mexico online
mexico drug stores pharmacies: Mexico pharmacy that ship to usa – mexico drug stores pharmacies
Доверьте нам свои желания, и мы превратим их в кулинарное произведение искусства https://vycypeo6kuhnyanazakaz.ru/!
mexican pharmacy: mexican northern doctors – mexican pharmacy
http://northern-doctors.org/# buying from online mexican pharmacy
Level up your game with insider tips and tricks for serious punters https://1xbetecricete1xbetei4.ru/.
mexican pharmacy: mexican pharmacy – mexico pharmacy
Experience the thrill of betting on cricket matches in the heart of the United Arab Emirates. Get ready to win big and cheer for your favorite teams https://1xbetecricete1xbetei4.ru/!
https://northern-doctors.org/# mexican drugstore online
purple pharmacy mexico price list: Mexico pharmacy that ship to usa – mexico drug stores pharmacies
Добро пожаловать в Кулинариум – место, где встречаются вкус и дизайн! Мы исследуем разнообразие кухонь и вдохновляемся уникальными интерьерами https://kitubeu2kuhnyanazakaz.ru/.
buying prescription drugs in mexico online: Mexico pharmacy that ship to usa – mexico drug stores pharmacies
https://northern-doctors.org/# mexican rx online
mexico drug stores pharmacies northern doctors pharmacy mexico pharmacies prescription drugs
Thanks for the tips you write about through this website. In addition, many young women that become pregnant tend not to even try to get medical care insurance because they fear they probably would not qualify. Although many states now require that insurers produce coverage irrespective of the pre-existing conditions. Premiums on these kind of guaranteed options are usually larger, but when thinking about the high cost of medical treatment it may be some sort of a safer approach to take to protect one’s financial potential.
reputable mexican pharmacies online: mexican pharmacy – mexican rx online
https://northern-doctors.org/# buying from online mexican pharmacy
buying prescription drugs in mexico online: northern doctors pharmacy – п»їbest mexican online pharmacies
At CricketBet, we bring the excitement of cricket betting to the vibrant landscape of the United Arab Emirates. Join us for a rollercoaster ride of predictions and wins https://1xbett1xbetc1xbetir4.ru/!
Our team is as passionate about cricket as you are, and we’re here to make your betting experience unforgettable https://1xbett1xbetc1xbetir4.ru/.
mexican drugstore online: mexican northern doctors – mexican pharmaceuticals online
http://northern-doctors.org/# mexican pharmaceuticals online
mexico drug stores pharmacies: mexican pharmacy northern doctors – mexico drug stores pharmacies
medication from mexico pharmacy northern doctors pharmacy best online pharmacies in mexico
Experience the thrill of betting on cricket matches in the heart of the United Arab Emirates. Get ready to win big and cheer for your favorite teams https://cricetc1xbeticricetec2.ru/!
Кухни про кухню! Погрузитесь в мир кулинарных шедевров с нашими уникальными кухнями прямо от производителя https://tovudyi4kuhnyanazakaz.ru/!
https://northern-doctors.org/# mexican mail order pharmacies
mexican pharmaceuticals online: reputable mexican pharmacies online – best online pharmacies in mexico
https://northern-doctors.org/# mexico pharmacy
mexican pharmacy: п»їbest mexican online pharmacies – mexico drug stores pharmacies
At CricketBet, we bring the excitement of cricket betting to the vibrant landscape of the United Arab Emirates. Join us for a rollercoaster ride of predictions and wins https://cricetc1xbetr1xbetcc2.ru/!
https://northern-doctors.org/# medication from mexico pharmacy
Welcome to the ultimate destination for cricket enthusiasts who want to spice up the game with some adrenaline-pumping bets https://cricetc1xbetr1xbetcc2.ru/!
mexican online pharmacies prescription drugs mexican pharmacy mexican border pharmacies shipping to usa
pharmacies in mexico that ship to usa: northern doctors – purple pharmacy mexico price list
mexican pharmacy: mexican pharmacy online – mexican rx online
https://northern-doctors.org/# mexican mail order pharmacies
mexico pharmacies prescription drugs: mexican northern doctors – medication from mexico pharmacy
Our team is as passionate about cricket as you are, and we’re here to make your betting experience unforgettable https://1xbetecricete1xbetei4.ru/.
pharmacies in mexico that ship to usa: mexican pharmacy online – medication from mexico pharmacy
mexico drug stores pharmacies: mexican pharmacy online – mexican pharmaceuticals online
https://northern-doctors.org/# purple pharmacy mexico price list
mexican mail order pharmacies: mexican northern doctors – mexico pharmacies prescription drugs
mexico drug stores pharmacies mexican pharmacy online purple pharmacy mexico price list
mexican rx online: mexican pharmacy northern doctors – mexico pharmacies prescription drugs
https://northern-doctors.org/# buying prescription drugs in mexico online
mexican drugstore online: mexican pharmacy – medication from mexico pharmacy
Experience the thrill of betting on cricket matches in the heart of the United Arab Emirates. Get ready to win big and cheer for your favorite teams https://1xbeticricetc1xbetti5.ru/!
Our team is as passionate about cricket as you are, and we’re here to make your betting experience unforgettable https://1xbeticricetc1xbetti5.ru/.
https://northern-doctors.org/# medication from mexico pharmacy
buying prescription drugs in mexico online: mexican pharmacy online – buying from online mexican pharmacy
purple pharmacy mexico price list: buying prescription drugs in mexico online – mexico pharmacy
One thing I’d prefer to say is the fact that before acquiring more computer system memory, take a look at the machine in which it will be installed. In the event the machine will be running Windows XP, for instance, the particular memory ceiling is 3.25GB. Putting in in excess of this would easily constitute a new waste. Make sure one’s mother board can handle an upgrade quantity, as well. Great blog post.
https://northern-doctors.org/# mexican drugstore online
mexico drug stores pharmacies: mexican pharmacy – mexican drugstore online
pharmacies in mexico that ship to usa mexican pharmacy online buying prescription drugs in mexico online
An fascinating dialogue is value comment. I feel that you need to write extra on this matter, it may not be a taboo topic but usually people are not sufficient to talk on such topics. To the next. Cheers
mexican border pharmacies shipping to usa: mexican northern doctors – mexican pharmacy
https://northern-doctors.org/# mexico pharmacy
п»їbest mexican online pharmacies: northern doctors – mexico drug stores pharmacies
http://northern-doctors.org/# mexico drug stores pharmacies
mexico pharmacies prescription drugs: mexican northern doctors – mexico drug stores pharmacies
mexican mail order pharmacies: mexican pharmacy online – mexico pharmacy
https://northern-doctors.org/# buying prescription drugs in mexico online
buying prescription drugs in mexico: mexican northern doctors – buying prescription drugs in mexico online
Level up your game with insider tips and tricks for serious punters https://cricetc1xbeticricetec2.ru/.
http://northern-doctors.org/# mexican border pharmacies shipping to usa
best online pharmacies in mexico: mexican pharmacy northern doctors – mexico drug stores pharmacies
Wonderful items from you, man. I have bear in mind your stuff previous to and you are just too great. I actually like what you have bought right here, really like what you are stating and the way in which through which you assert it. You’re making it enjoyable and you still take care of to keep it sensible. I can not wait to read much more from you. That is actually a tremendous web site.
mexican border pharmacies shipping to usa cmq mexican pharmacy online purple pharmacy mexico price list
mexican pharmacy: mexico pharmacy – mexican online pharmacies prescription drugs
mexican pharmaceuticals online
https://cmqpharma.com/# mexican rx online
mexico pharmacies prescription drugs
medicine in mexico pharmacies mexican pharmacy medicine in mexico pharmacies
mexican online pharmacies prescription drugs mexican pharmacy online pharmacies in mexico that ship to usa
pharmacies in mexico that ship to usa mexican online pharmacy mexico pharmacies prescription drugs
mexican rx online cmq mexican pharmacy online pharmacies in mexico that ship to usa
п»їbest mexican online pharmacies cmqpharma.com buying prescription drugs in mexico online
One more issue is that video games are generally serious as the name indicated with the most important focus on mastering rather than leisure. Although, it has an entertainment element to keep children engaged, just about every game is usually designed to work with a specific group of skills or curriculum, such as numbers or technology. Thanks for your write-up.
I would also love to add that if you do not currently have an insurance policy otherwise you do not participate in any group insurance, chances are you’ll well reap the benefits of seeking the aid of a health agent. Self-employed or those that have medical conditions ordinarily seek the help of one health insurance brokerage. Thanks for your writing.
mexican online pharmacies prescription drugs online mexican pharmacy mexican drugstore online
Want to improve your SEO rankings and save time? Our premium databases for XRumer and GSA Search Engine Ranker are just what you need!
What do our databases include?
• Active links: Get access to constantly updated lists of active links from profiles, posts, forums, guestbooks, blogs, and more. No more wasting time on dead links!
• Verified and identified links: Our premium databases for GSA Search Engine Ranker include verified and identified links, categorized by search engines. This means you get the highest quality links that will help you rank higher.
• Monthly updates: All of our databases are updated monthly to ensure you have the most fresh and effective links.
Choose the right option for you:
• XRumer premium database:
o Premium database with free updates: $119
o Premium database without updates: $38
• Fresh XRumer Database:
o Fresh database with free updates: $94
o Fresh database without updates: $25
• GSA Search Engine Ranker Verified Links:
o GSA Search Engine Ranker activation key: $65 (includes database)
o Fresh database with free updates: $119
o Fresh database without updates: $38
Don’t waste time on outdated or inactive links. Invest in our premium databases and start seeing results today!
Order now!
P.S. By purchasing GSA Search Engine Ranker from us, you get a high-quality product at a competitive price. Save your resources and start improving your SEO rankings today!
To contact us, write to telegram https://t.me/DropDeadStudio
Today, I went to the beachfront with my kids. I found a sea shell and gave it to my 4 year old daughter and said “You can hear the ocean if you put this to your ear.” She placed the shell to her ear and screamed. There was a hermit crab inside and it pinched her ear. She never wants to go back! LoL I know this is totally off topic but I had to tell someone!
Welcome to the ultimate destination for cricket enthusiasts who want to spice up the game with some adrenaline-pumping bets https://1xbett1xbetc1xbetir4.ru/!
Wow, marvelous blog layout! How lengthy have you ever been running a blog for? you made blogging look easy. The overall glance of your website is excellent, as neatly as the content material!
At CricketBet, we bring the excitement of cricket betting to the vibrant landscape of the United Arab Emirates. Join us for a rollercoaster ride of predictions and wins https://cricetc1xbeticricetec2.ru/!
Experience the thrill of betting on cricket matches in the heart of the United Arab Emirates. Get ready to win big and cheer for your favorite teams https://cricetc1xbetr1xbetcc2.ru/!
purple pharmacy mexico price list cmq pharma mexican pharmacy medicine in mexico pharmacies
http://cmqpharma.com/# mexican rx online
mexico drug stores pharmacies
I was recommended this web site by my cousin. I am not sure whether this post is written by him as nobody else know such detailed about my trouble. You’re incredible! Thanks!
I am really enjoying the theme/design of your weblog. Do you ever run into any web browser compatibility issues? A number of my blog audience have complained about my site not working correctly in Explorer but looks great in Firefox. Do you have any solutions to help fix this issue?
Wow! I’m in awe of the author’s writing skills and talent to convey complex concepts in a straightforward and concise manner. This article is a true gem that deserves all the accolades it can get. Thank you so much, author, for sharing your expertise and offering us with such a precious asset. I’m truly appreciative!
This article is a breath of fresh air! The author’s unique perspective and perceptive analysis have made this a truly engrossing read. I’m appreciative for the effort she has put into producing such an educational and mind-stimulating piece. Thank you, author, for sharing your wisdom and stimulating meaningful discussions through your brilliant writing!
Hi my friend! I want to say that this post is awesome, nice written and include almost all important infos. I?d like to see more posts like this.
I do not even know how I ended up here, but I thought this post was good. I do not know who you are but definitely you are going to a famous blogger if you are not already 😉 Cheers!
I think this is one of the most significant info for me. And i am glad reading your article. But wanna remark on some general things, The web site style is great, the articles is really nice : D. Good job, cheers
Hey there! Would you mind if I share your blog with my zynga group? There’s a lot of people that I think would really enjoy your content. Please let me know. Cheers
Greetings! I’ve been reading your site for some time now and finally got the courage to go ahead and give you a shout out from Houston Texas! Just wanted to say keep up the great work!
mexican mail order pharmacies
https://cmqpharma.online/# best online pharmacies in mexico
pharmacies in mexico that ship to usa
mexican pharmaceuticals online: cmq pharma – best online pharmacies in mexico
Usually I don’t read article on blogs, but I would like to say that this write-up very forced me to try and do it! Your writing style has been amazed me. Thanks, quite nice article.
Nice post. I learn something more challenging on different blogs everyday. It will always be stimulating to read content from other writers and practice a little something from their store. I’d prefer to use some with the content on my blog whether you don’t mind. Natually I’ll give you a link on your web blog. Thanks for sharing.
Thanks for your publication on this blog site. From my very own experience, many times softening upward a photograph may possibly provide the professional photographer with an amount of an artsy flare. More often than not however, that soft cloud isn’t what precisely you had in mind and can quite often spoil a normally good picture, especially if you thinking about enlarging it.
mexican online pharmacies prescription drugs: mexican pharmacy online – mexico pharmacies prescription drugs
best online pharmacies in mexico
http://cmqpharma.com/# mexican online pharmacies prescription drugs
buying from online mexican pharmacy
Thanks for a marvelous posting! I actually enjoyed reading it, you could be a great author.I will make certain to bookmark your blog and may come back in the foreseeable future. I want to encourage you to definitely continue your great job, have a nice weekend!
What is Gluco6? Gluco6 is a revolutionary dietary supplement designed to help individuals manage their blood sugar levels naturally.
One other issue is when you are in a predicament where you will not have a cosigner then you may actually want to try to wear out all of your financial aid options. You will find many funds and other scholarship grants that will present you with finances to help you with school expenses. Thanks for the post.
Thank you, I have recently been looking for details about this topic for ages and yours is the best I’ve found so far.
My spouse and I absolutely love your blog and find a lot of your post’s to be what precisely I’m looking for. can you offer guest writers to write content to suit your needs? I wouldn’t mind publishing a post or elaborating on a lot of the subjects you write regarding here. Again, awesome web site!
I’ve read a few just right stuff here. Definitely price bookmarking for revisiting. I surprise how a lot attempt you set to create this sort of fantastic informative site.
When I originally commented I clicked the “Notify me when new comments are added” checkbox and now each time a comment is added I get three e-mails with the same comment. Is there any way you can remove people from that service? Cheers!
I?¦ll immediately grasp your rss feed as I can’t to find your email subscription hyperlink or newsletter service. Do you’ve any? Kindly let me recognise so that I could subscribe. Thanks.
I simply could not leave your site prior to suggesting that I actually enjoyed the usual information an individual supply in your guests? Is going to be back incessantly to check out new posts
Hi my loved one! I want to say that this article is awesome, great written and include approximately all important infos. I?¦d like to look more posts like this .
I want to show my respect for your kindness for men and women who really want assistance with this particular theme. Your very own commitment to passing the solution throughout had been extraordinarily powerful and has specifically allowed somebody just like me to get to their objectives. Your warm and helpful publication signifies so much to me and somewhat more to my fellow workers. Thank you; from everyone of us.
Level up your game with insider tips and tricks for serious punters https://1xbeticricetc1xbetti5.ru/.
Sex
Pretty nice post. I just stumbled upon your blog and wished to say that I have really enjoyed surfing around your blog posts. After all I’ll be subscribing to your rss feed and I hope you write again very soon!
Porn site
Heya i’m for the first time here. I came across this board and I find It really useful & it helped me out a lot. I hope to give something back and aid others like you helped me.
Sex
Pornstar
Viagra
Sex
Buy Drugs
Thanx for the effort, keep up the good work Great work, I am going to start a small Blog Engine course work using your site I hope you enjoy blogging with the popular BlogEngine.net.Thethoughts you express are really awesome. Hope you will right some more posts.
I am glad to be a visitant of this gross web site! , appreciate it for this rare information! .
Viagra
Porn site
Scam
Scam
Sex
Porn site
Scam
After I originally commented I clicked the -Notify me when new feedback are added- checkbox and now every time a remark is added I get four emails with the identical comment. Is there any approach you possibly can remove me from that service? Thanks!
Porn
Pornstar
Porn
Only wanna input that you have a very decent internet site, I like the layout it actually stands out.
Buy Drugs
Porn
Viagra
Viagra
Buy Drugs
I’m extremely impressed with your writing skills as well as with the layout on your blog. Is this a paid theme or did you modify it yourself? Anyway keep up the excellent quality writing, it is rare to see a great blog like this one today..
Porn
Porn site
Sex
Porn
Pornstar
Viagra
Pornstar
Viagra
Porn
Porn site
Buy Drugs
Buy Drugs
Awsome website! I am loving it!! Will be back later to read some more. I am taking your feeds also
Sex
Sex
Pornstar
Porn
Sex
Sex
Buy Drugs
canadapharmacyonline legit: safe canadian pharmacy – safe canadian pharmacies
http://indiapharmast.com/# india pharmacy mail order
canadian pharmacy online canadian pharmacy 24 canadian pharmacy meds reviews
Porn site
pharmacy website india: buy prescription drugs from india – indian pharmacy paypal
Porn
mexican rx online: mexican rx online – mexican rx online
Buy Drugs
Buy Drugs
Scam
Porn
Its like you read my mind You appear to know so much about this like you wrote the book in it or something I think that you can do with a few pics to drive the message home a little bit but instead of that this is excellent blog A fantastic read Ill certainly be back
world pharmacy india: online pharmacy india – pharmacy website india
Porn site
indian pharmacy paypal online pharmacy india online pharmacy india
http://foruspharma.com/# mexican border pharmacies shipping to usa
I like what you guys at kitchen remodeling lansing are up to Such smart professional design and craftsmanship! Keep up the superb works guys I’ve incorporated you guys to my contacts I think by my passing along kitchen lansing remodeling info it will improve the value of my credibility
cheapest online pharmacy india: mail order pharmacy india – best online pharmacy india
mexican pharmaceuticals online: best online pharmacies in mexico – mexico pharmacy
Porn
buy drugs from canada safe online pharmacies in canada canadian pharmacy uk delivery
best rated canadian pharmacy: canadian mail order pharmacy – reputable canadian online pharmacies
http://foruspharma.com/# purple pharmacy mexico price list
reliable canadian pharmacy: online canadian pharmacy reviews – online canadian pharmacy
medication canadian pharmacy: maple leaf pharmacy in canada – canadapharmacyonline
Pornstar
Porn site
Sex
Sex
mexican drugstore online: best online pharmacies in mexico – buying prescription drugs in mexico
Buy Drugs
Porn
Buy Drugs
Porn
canadapharmacyonline com: canadian pharmacy – pharmacy canadian
Sex
Sex
Paxlovid over the counter: paxlovid price – paxlovid cost without insurance
Pornstar
http://amoxildelivery.pro/# generic amoxicillin online
Scam
Наши кухни – это не просто мебель это произведения искусства которые вдохновляют на кулинарные подвиги https://xehyroo5kuhnyanazakaz.ru/
У нас вы найдете самые необычные идеи для вашей кухни которые заставят вас влюбиться в готовку снова https://kitubeu2kuhnyanazakaz.ru/
Наши кухни – это не просто мебель это искусство которое оживляет ваш дом https://tovudyi4kuhnyanazakaz.ru/.
Кухни про кухню Погрузитесь в мир кулинарных шедевров с нашими уникальными кухнями прямо от производителя https://tovudyi4kuhnyanazakaz.ru/
Присоединяйтесь к нам и откройте новый уровень кулинарного опыта https://guzywia4kuhnyanazakaz.ru/
Откройте мир волшебства в каждой кухне от фабрики КухниФаб. Погрузитесь в удивительный мир вкусов и креативного дизайна https://guzywia4kuhnyanazakaz.ru/
Мы – команда талантливых дизайнеров и мастеров готовых воплотить в жизнь любую кухню вашей мечты https://vycypeo6kuhnyanazakaz.ru/.
Some times its a pain in the ass to read what website owners wrote but this website is very user genial! .
Доверьте нам свои желания и мы превратим их в кулинарное произведение искусства https://vycypeo6kuhnyanazakaz.ru/
Присоединяйтесь к нам и создайте кухню вашей мечты https://xehyroo5kuhnyanazakaz.ru/
Превратите свою кухню в шедевр https://xehyroo5kuhnyanazakaz.ru/
http://clomiddelivery.pro/# how can i get clomid
Buy Drugs
Доверьте нам создание кухни которая станет сердцем вашего пространства https://tovudyi4kuhnyanazakaz.ru/.
Мы – команда страстных кулинаров и дизайнеров создающая кухни мечты для вас https://tovudyi4kuhnyanazakaz.ru/
Buy Drugs
can i get generic clomid without insurance: clomid medication – can you get cheap clomid without a prescription
Мы – команда страстных энтузиастов создающих кухни которые превращают готовку в волшебство https://guzywia4kuhnyanazakaz.ru/.
My spouse and I absolutely love your blog and find almost all of your post’s to be exactly I’m looking for. can you offer guest writers to write content for yourself? I wouldn’t mind writing a post or elaborating on many of the subjects you write in relation to here. Again, awesome web log!
Porn
Buy Drugs
Viagra
Наши кухни не просто мебель это произведения искусства наполненные вдохновением и инновациями https://guzywia4kuhnyanazakaz.ru/.
У нас нет предела совершенству Мы создаем гарнитуры которые будут радовать вас каждый день https://vycypeo6kuhnyanazakaz.ru/.
https://ciprodelivery.pro/# buy cipro online canada
Уникальные кухонные шедевры ждут вас https://vycypeo6kuhnyanazakaz.ru/
Sex
Sex
Viagra
Sex
Погрузитесь в мир кулинарных искусств где каждая кухня – это отражение стиля и вкуса https://kitubeu2kuhnyanazakaz.ru/.
Sex
http://paxloviddelivery.pro/# paxlovid for sale
buy cipro no rx: buy cipro without rx – cipro ciprofloxacin
You really make it seem really easy along with your presentation however I find this matter to be really something which I feel I’d never understand. It sort of feels too complicated and extremely vast for me. I am taking a look ahead in your next submit, I will attempt to get the grasp of it!
Pornstar
Добро пожаловать в Кулинариум – место где встречаются вкус и дизайн Мы исследуем разнообразие кухонь и вдохновляемся уникальными интерьерами https://kitubeu2kuhnyanazakaz.ru/.
Viagra
http://amoxildelivery.pro/# buy amoxicillin without prescription
Мы – команда талантливых дизайнеров воплощаем в жизнь ваши самые дерзкие кулинарные фантазии https://xehyroo5kuhnyanazakaz.ru/.
Viagra
Scam
Scam
I do enjoy the manner in which you have presented this specific issue plus it does provide us a lot of fodder for thought. However, coming from everything that I have seen, I just hope when the commentary stack on that people continue to be on issue and don’t get started on a soap box of some other news of the day. All the same, thank you for this exceptional point and whilst I do not agree with the idea in totality, I regard your perspective.
amoxicillin without a doctors prescription: order amoxicillin no prescription – amoxicillin for sale
Buy Drugs
Scam
Viagra
I love it when people come together and share opinions, great blog, keep it up.
https://paxloviddelivery.pro/# Paxlovid buy online
Pornstar
Porn site
http://doxycyclinedelivery.pro/# buy doxycycline
buy cipro online usa: antibiotics cipro – ciprofloxacin generic
Please let me know if you’re looking for a writer for your weblog. You have some really good articles and I believe I would be a good asset. If you ever want to take some of the load off, I’d absolutely love to write some material for your blog in exchange for a link back to mine. Please shoot me an e-mail if interested. Thanks!
http://amoxildelivery.pro/# order amoxicillin online uk
Porn
Way cool, some valid points! I appreciate you making this article available, the rest of the site is also high quality. Have a fun.
Porn site
https://paxloviddelivery.pro/# paxlovid price
amoxicillin 500 mg without a prescription: amoxicillin no prescipion – amoxicillin without prescription
Porn site
Pornstar
Scam
Scam
Viagra
Attractive section of content. I just stumbled upon your web site and in accession capital to assert that I get actually enjoyed account your blog posts. Anyway I’ll be subscribing to your augment and even I achievement you access consistently quickly.
Pornstar
Sex
Hello. splendid job. I did not anticipate this. This is a fantastic story. Thanks!
Viagra
Buy Drugs
amoxicillin 250 mg: amoxicillin 800 mg price – cost of amoxicillin 30 capsules
Buy Drugs
Sex
Magnificent website. Plenty of useful info here. I¦m sending it to a few friends ans additionally sharing in delicious. And certainly, thank you for your sweat!
You are my breathing in, I possess few web logs and infrequently run out from to post .
Porn site
Porn
Pornstar
Scam
Buy Drugs
Viagra
Oh my goodness! a tremendous article dude. Thanks Nonetheless I am experiencing challenge with ur rss . Don’t know why Unable to subscribe to it. Is there anybody getting identical rss drawback? Anyone who knows kindly respond. Thnkx
Very interesting details you have noted, thankyou for posting.
Pornstar
Some genuinely good posts on this web site, thanks for contribution.
Pornstar
Porn
Some really nice stuff on this site, I like it.
Porn site
Viagra
Porn
I was very pleased to find this web-site.I wanted to thanks for your time for this wonderful read!! I definitely enjoying every little bit of it and I have you bookmarked to check out new stuff you blog post.
Советы по предотвращению дубликатов номеров Полезные советы по предотвращению дубликатов номеров Опасности использования одинаковых номеров Полезные рекомендации по пронумерации документов Зачем важно избегать дубликатов номеров Частые причины появления одинаковых номеров Образцы нумерации для предотвращения повторений Как легко обнаружить дубликаты номеров Как исправить повторения номеров Шаги по устранению дубликатов номеров Полновесные рекомендации по избежанию дубликатов номеров в публикациях Рекомендации по нумерации для предотвращения повторений Как использовать номера умнее чтобы избежать дубликатов Как документировать процесс нумерации для предотвращения дубликатов номеров Опасности повторения номеров в документах Способы использования технологий для предотвращения дубликатов Как избежать оштрафования за дубликаты номеров в документах Способы обезопасить свои документы от повторений номеров Как стать мастером в избежании одинаковых номеров. гос номер изготовление гос номер изготовление .
Как избежать дубликатов номеров в своих документах Полезные советы по предотвращению дубликатов номеров Опасности использования одинаковых номеров Полезные рекомендации по пронумерации документов Почему следует избегать одинаковых номеров в тексте Какие ошибки нумерации могут привести к проблемам Примеры правильной нумерации Как легко обнаружить дубликаты номеров Полезные рекомендации по исправлению дубликатов номеров Как реагировать на повторения номеров Полновесные рекомендации по избежанию дубликатов номеров в публикациях Рекомендации по нумерации для предотвращения повторений Техники использования уникальных номеров Как документировать процесс нумерации для предотвращения дубликатов номеров Почему каждый должен бороться с дубликатами номеров Программы для поиска повторений номеров Как избежать оштрафования за дубликаты номеров в документах Способы обезопасить свои документы от повторений номеров Как стать мастером в избежании одинаковых номеров. изготовление дубликатов https://dublikat-nomerov.ru/ .